| Structure title | Structure of a yeast 48S-AUC preinitiation complex in closed conformation |
|---|---|
| Method | ELECTRON MICROSCOPY |
| Resolution | 3.35 Å |
| Source organism | Kluyveromyces lactis NRRL Y-1140 |
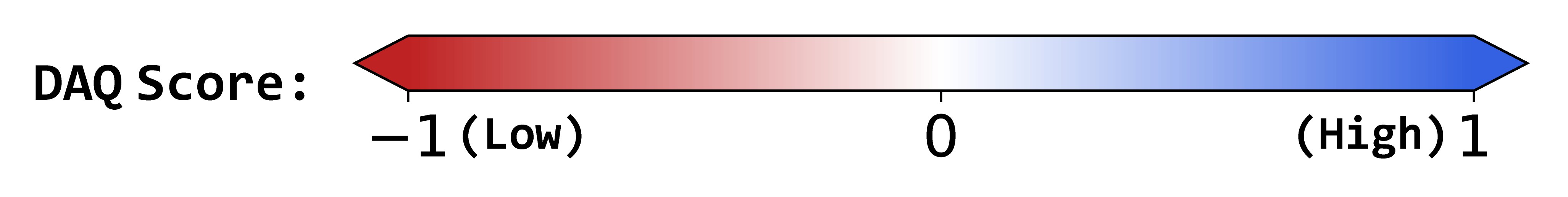
DAQ Score Table
0.12
1.31
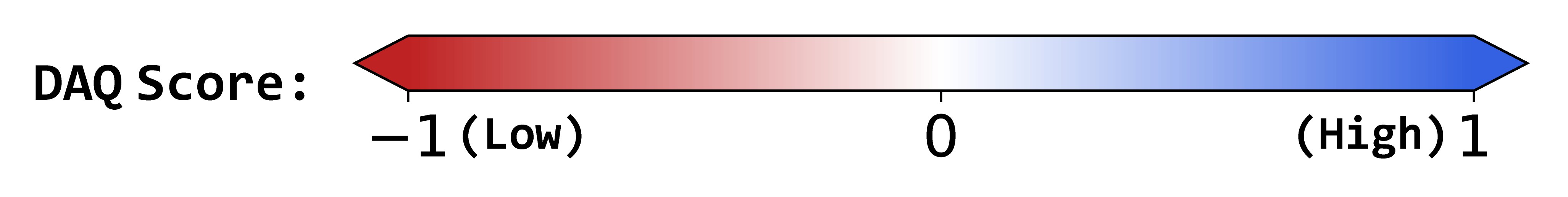
DAQ Score Table
0
1.09
DAQ Score Table
-0.15
0.88