| Structure title | NTD of GluA2 in complex with CNIH3 - with antagonist ZK200775 - in asymmetric global conformation |
|---|---|
| Method | ELECTRON MICROSCOPY |
| Resolution | 3.12 Å |
| Source organism | Rattus norvegicus |
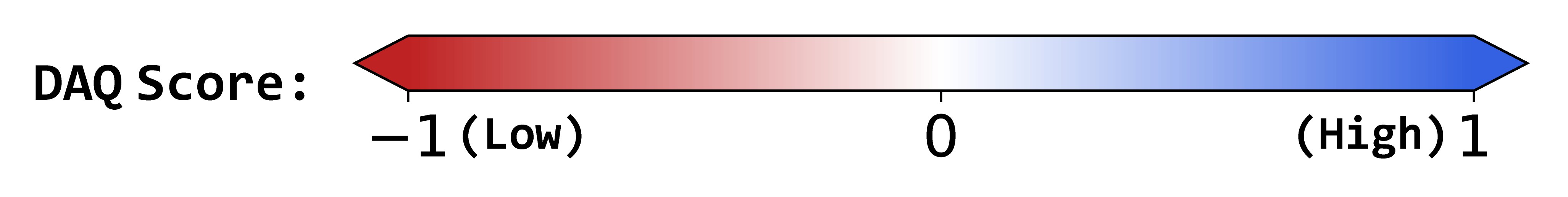
DAQ Score Table
0.29
1.46
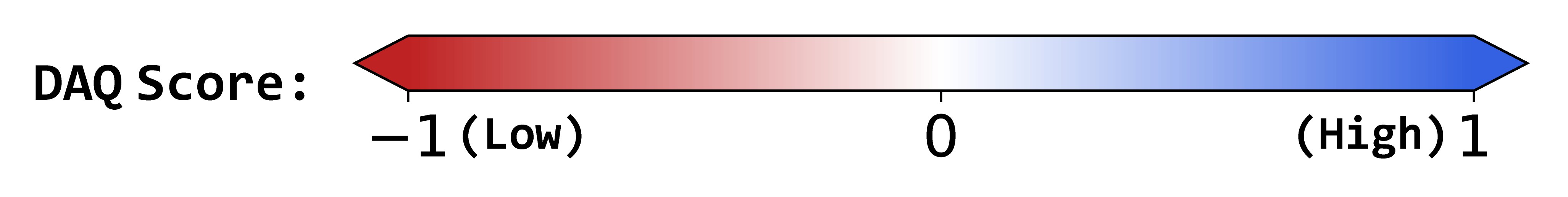
DAQ Score Table
0.33
1.25