| Structure title | Cryo-EM structure of the Arabidopsis thaliana photosystem I(PSI-LHCII-ST2) |
|---|---|
| Method | ELECTRON MICROSCOPY |
| Resolution | 2.79 Å |
| Source organism | Arabidopsis thaliana |
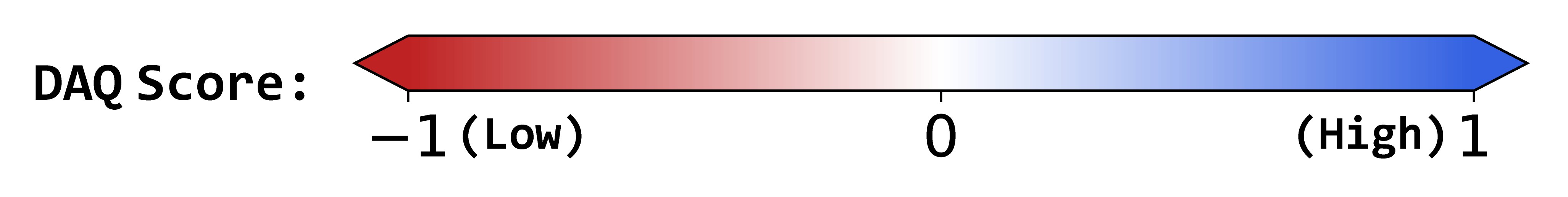
DAQ Score Table
0.37
1.55
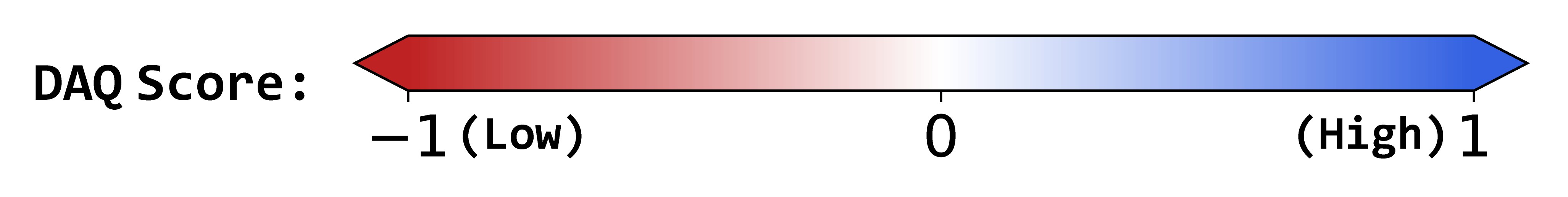
DAQ Score Table
0.30
1.21
DAQ Score Table
-0.72
1.03